Call for Abstracts for the 17th Annual Bioinformatics Open Source Conference (BOSC 2016), a Special Interest Group (SIG) of ISMB 2016.
- Dates: July 8-9, 2016
- Location: Orlando, FL
- Web site: /wiki/BOSC_2016
- Email: bosc@open-bio.org
- BOSC announcements mailing list: http://lists.open-bio.org/mailman/listinfo/bosc-announce
- Twitter: @OBF_BOSC and @OBF_News
Important Dates:
- Call for one-page abstracts opens: March 1, 2016
- Abstract submission deadline: April 1, 2016 - extended to Monday 4 April 2016
- Travel fellowship application deadline: April 15, 2016
- Authors notified: May 6, 2016
- Codefest 2016: July 6-7, 2016, Orlando, FL (confirming venue)
- BOSC 2016: July 8-9, 2016, Orlando, FL
- ISMB 2016: July 8-12, 2016, Orlando, FL
The Bioinformatics Open Source Conference (BOSC) is run as a two-day meeting before the annual ISMB conference. It is organized by the Open Bioinformatics Foundation (OBF), a non-profit group dedicated to promoting the practice and philosophy of open source software development and open science within the biological research community. BOSC offers a focused environment for developers and users to interact and share ideas about standards; software development practices; practical techniques for solving bioinformatics problems; and approaches that promote open science and sharing of data, results and software.
[Read More]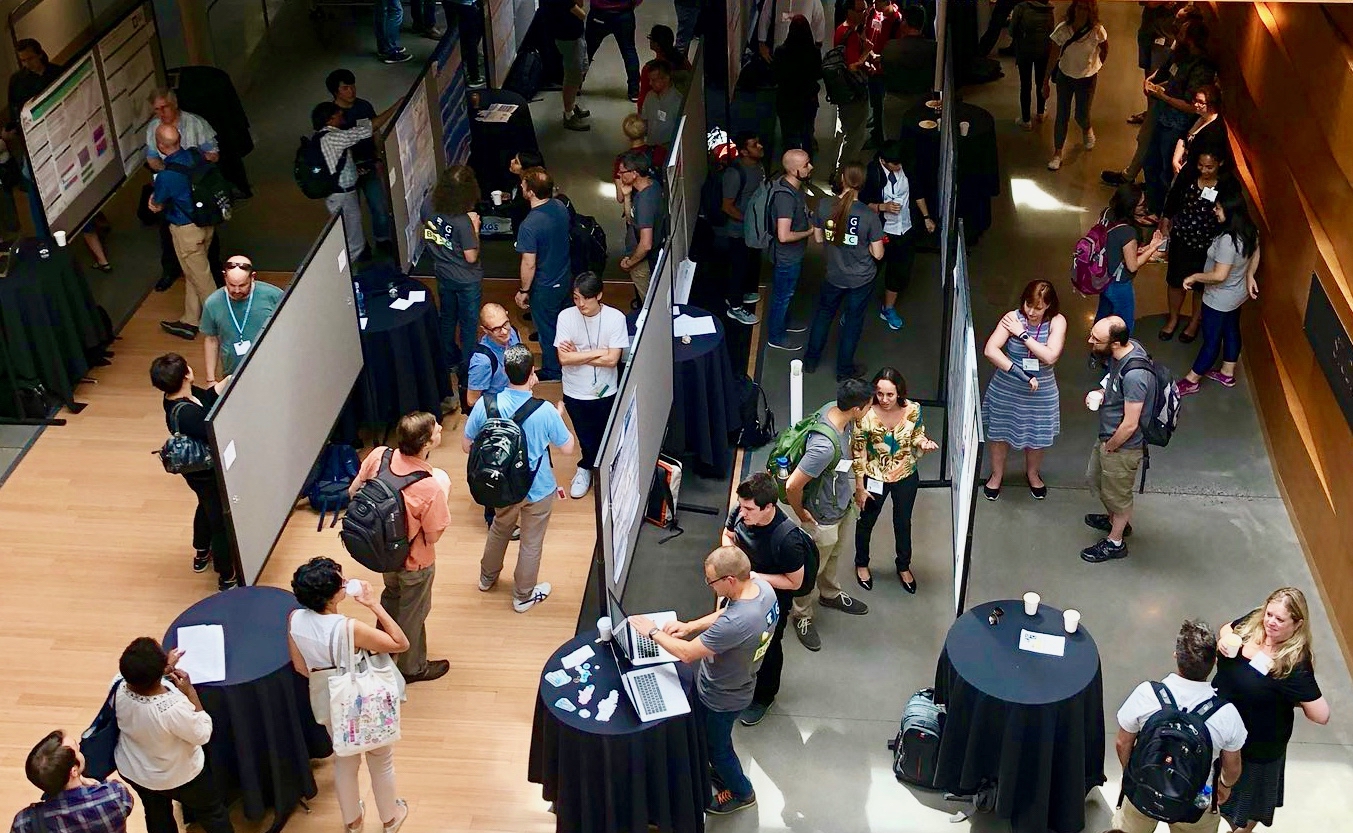

![[BOSC Logo]](/w/images/b/b0/Pear.png)
![[ISMB 2016 logo]](https://news.obf.io/wp-content/uploads/2016/03/ismb2016.png)
