Upgraded MW install for some bugfixes and security fixes.
About the OBF
The Open Bioinformatics Foundation (OBF) is a non-profit, volunteer-run group that promotes open source software development and Open Science within the biological research community. Membership in the OBF is free and open to anyone who wants to help promote open source or open science in a biological field.
OBF runs the annual Bioinformatics Open Source Conference (BOSC).
BOSC 2025 took place July 21-22, 2025, in Liverpool, UK (as part of ISMB/ECCB 2025). BOSC 2026 will be part of ISMB 2026 in Washington, DC.
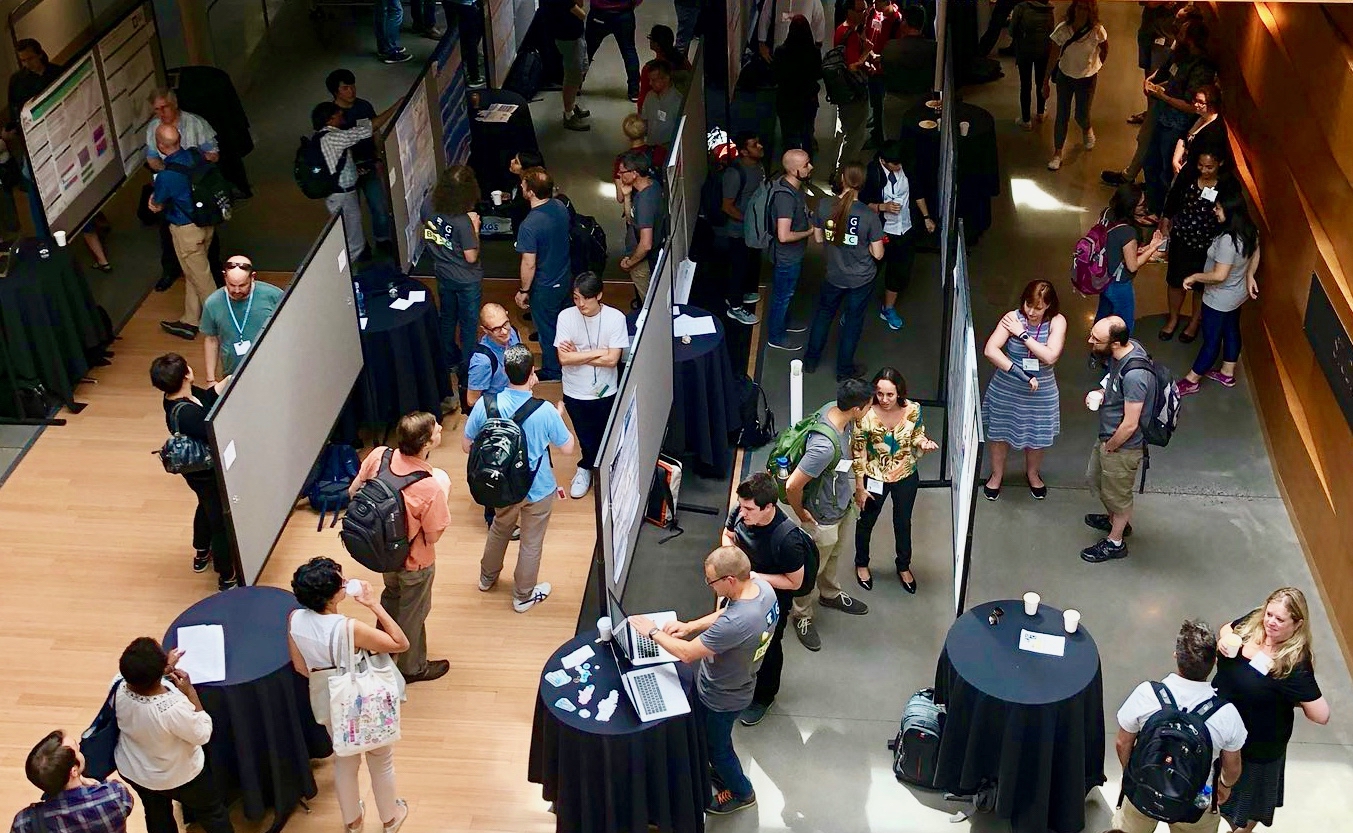
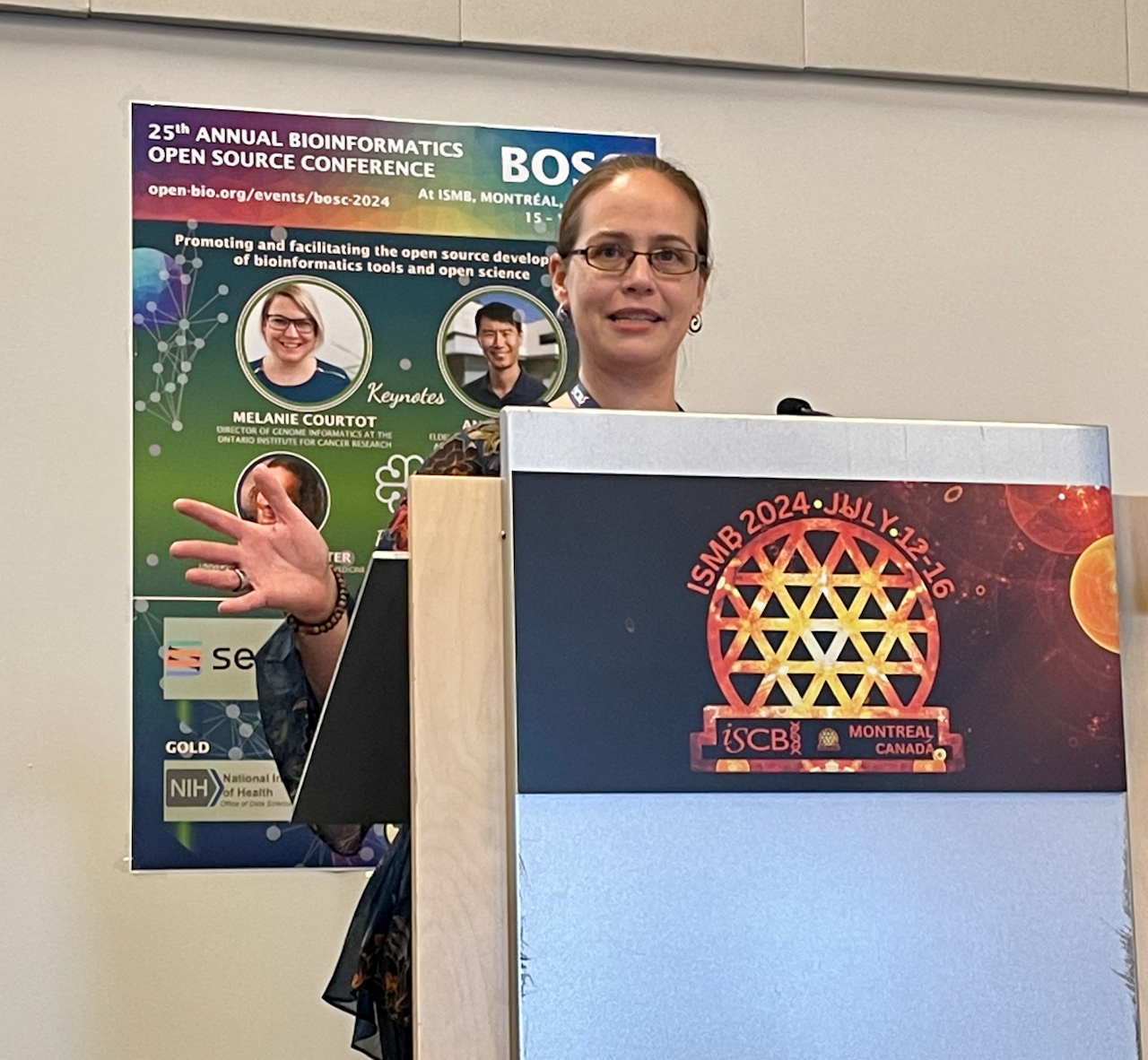


OBF Event Awards
The OBF Event Fellowship program aims to increase diverse participation at events promoting open source bioinformatics software development and open science in the biological research community.
bioperl-network added to Pdoc
Bioperl Network package documentation added to the Pdoc site and is automaticly updated daily.
Mediawiki upgraded
Upgraded BioPerl mediawiki to 1.6.6.
Modware: a BioPerl based API for Chado
We are announcing a new Sourceforge Project called Modware. It is an object-oriented API written in Perl that creates BioPerl object representations of biological features stored in a Chado database. It basically creates a Bio::Seq object for chromosomes in Chado and creates Bio::SeqFeature::Gene objects for protein coding transcripts stored in Chado. Things like contigs are represented as Bio::SeqFeature::Generic objects. We also provide many methods for manipulating these objects once they are in memory.
[Read More]Deobfuscator interface now available
I’m glad to announce the availability of the Deobfuscator interface at the BioPerl website. You can use it at the following URL:
http://bioperl.org/cgi-bin/deob_interface.cgi
Many thanks to Laura Kavanaugh and David Messina for this great contribution to the BioPerl project!
ListSummaries for April 26-May 9
ListSummaries for April 26-May 9 are up at the usual place:
http://www.bioperl.org/wiki/Mailing_list_summaries
Direct link:
http://www.bioperl.org/wiki/ListSummary:April_26-May_9%2C2006
It’s a bit of a hurried one so don’t be surprised to find a few spelling errors here and there. I’m getting ready for a conference in a couple weeks so I may be off the radar a bit here and there. The next ListSummary won’t be posted until May 26. Enjoy!
BioPerl-run in FreeBSD
It’s my great pleasure to announce the availability of the BioPerl-run packages (stable & developer releases) for the FreeBSD operating system.
For instructions on how to install BioPerl ports in FreeBSD, please take a look into the Getting Bioperl section of the BioPerl Wiki.
RSS feeds
I’ve added a page about RSS feeds in BioPerl. These include links to CVS commits as a RSS feed. This a bit of a hack using cvs2rss and cvs2cl and I have hardcoded it to show the last 30 commits only.
In addition RSS news is now embedded on the main BioPerl and Tracking CVS commits webpages to make for better interlinking between the news and wiki site ( you might even be reading this there).
[Read More]Bioperl ListSummaries for April 12-25
The newest summary of the BioPerl mailing lists has been posted to the wiki:
Post gripes, harrassments, and faint praises at the regular places.
BioPerl Mailing List Summaries
The first of a biweekly summary of BioPerl mailing list summaries has been posted to the wiki:
These will likely be archived on the wiki but may be moved to a more suitable location in the future (maybe to this blog?).