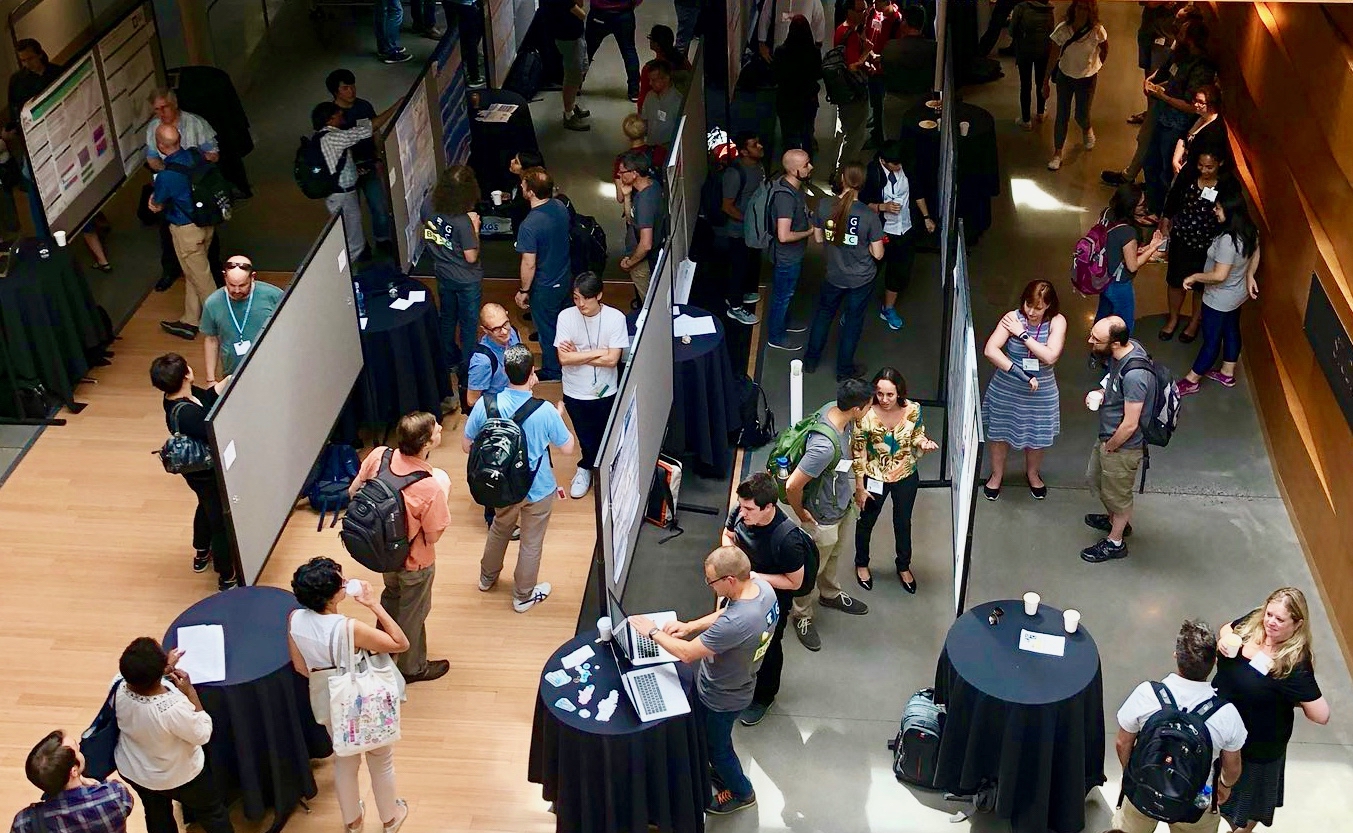
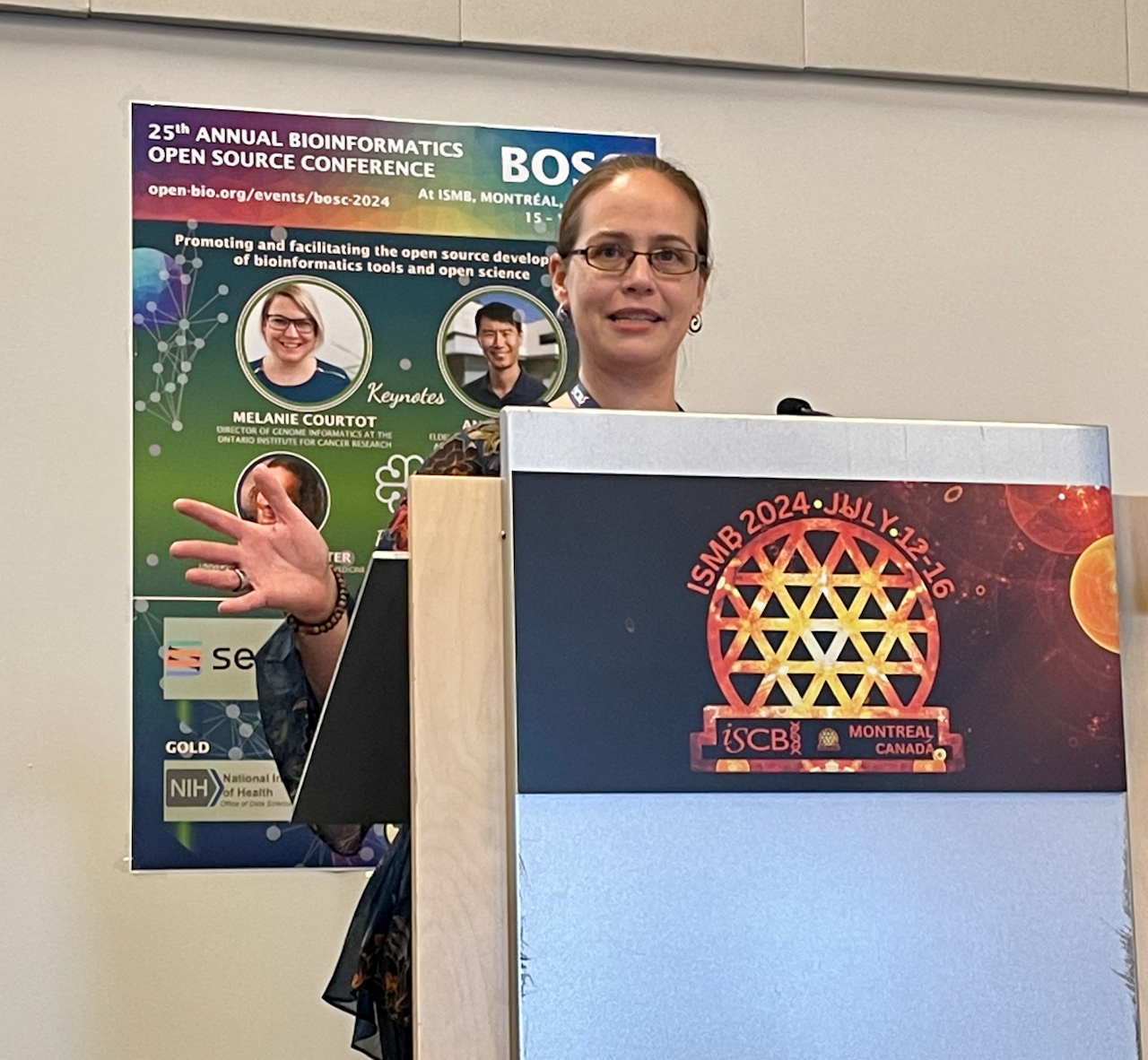
Venue: Renaissance Center in Detroit, on the first floor next to the Starbucks. Date: June 23, 2005, 16:15.
Call to order:
- Board members present: Chris, Hilmar, Ewan, Andrew
- Others: Jason Stajich
Chris / Treasurer’s Report
- Review of the accounts / passing the statements around
- Summary of response to the fraud attempt
- Request to activate the debit card. Granted.
- The account got a $9,000 deposit but we don’t know who it comes from. Chris will investigate.
- We have not yet received our money from the Glasgow BOSC. Chris will follow up on that.
- Chris will pull the current credit report for Open Bio.
- Develop a succession / “hit by the bus” plan for hardware and people. Eg, can we get access to the account if Chris decides to be a beach bum in the S. Pacific?
- We did not pay Delaware to renew our corporate registration. Chris will look into getting the right person to fix that and pay any penalties.
- Some discussion about letting Chris make minor (<$1,000) payments without a counter-signature. Hilmar pointed out that’s in the new bylaws
New Bylaws, Membership
Hilmar presentation of the new OBF bylaws. We reviewed it and made comments. Some time later he finished.
[Read More]