About the OBF
The Open Bioinformatics Foundation (OBF) is a non-profit, volunteer-run group that promotes open source software development and Open Science within the biological research community. Membership in the OBF is free and open to anyone who wants to help promote open source or open science in a biological field.
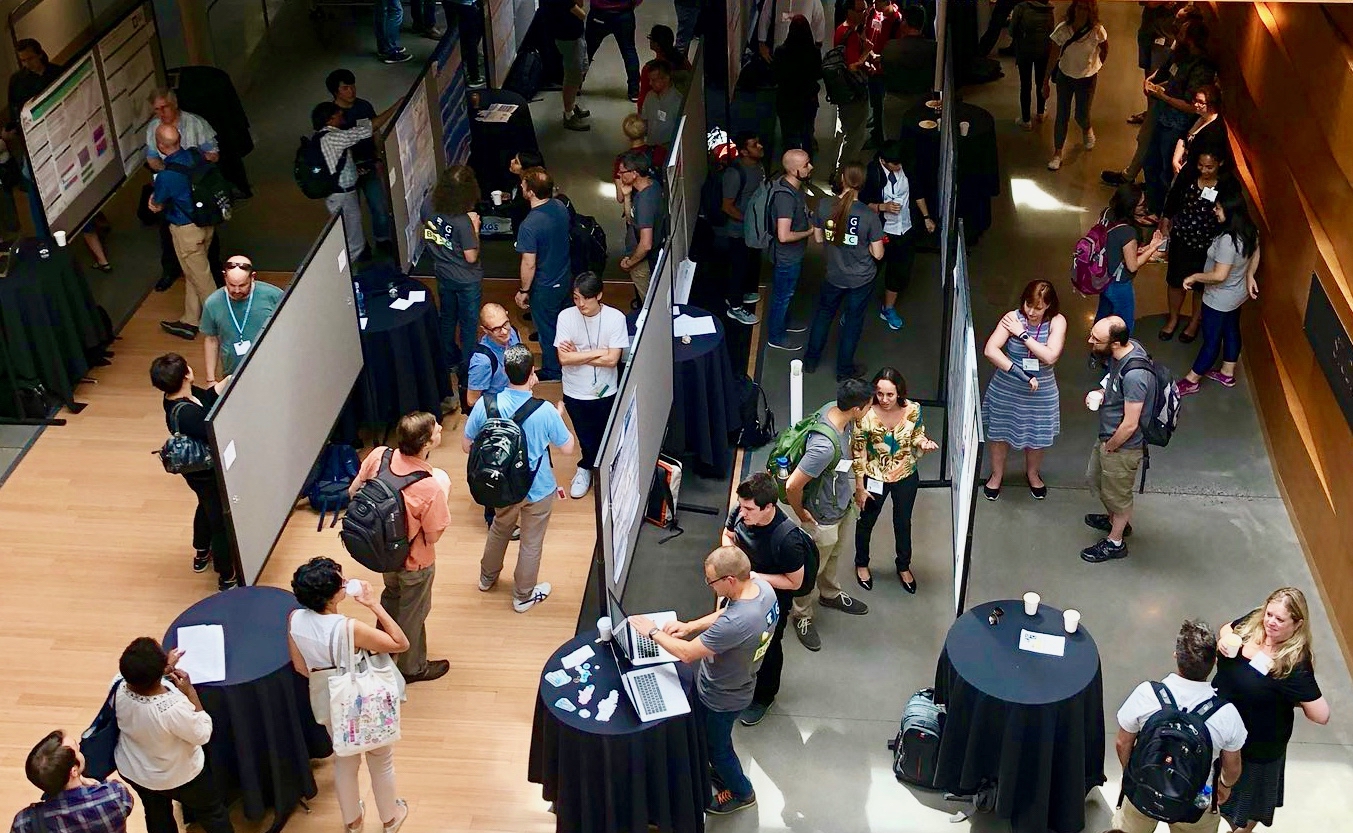
OBF runs the annual Bioinformatics Open Source Conference (BOSC).
BOSC 2025 took place July 21-22, 2025, in Liverpool, UK (as part of ISMB/ECCB 2025). BOSC 2026 will be part of ISMB 2026 in Washington, DC.
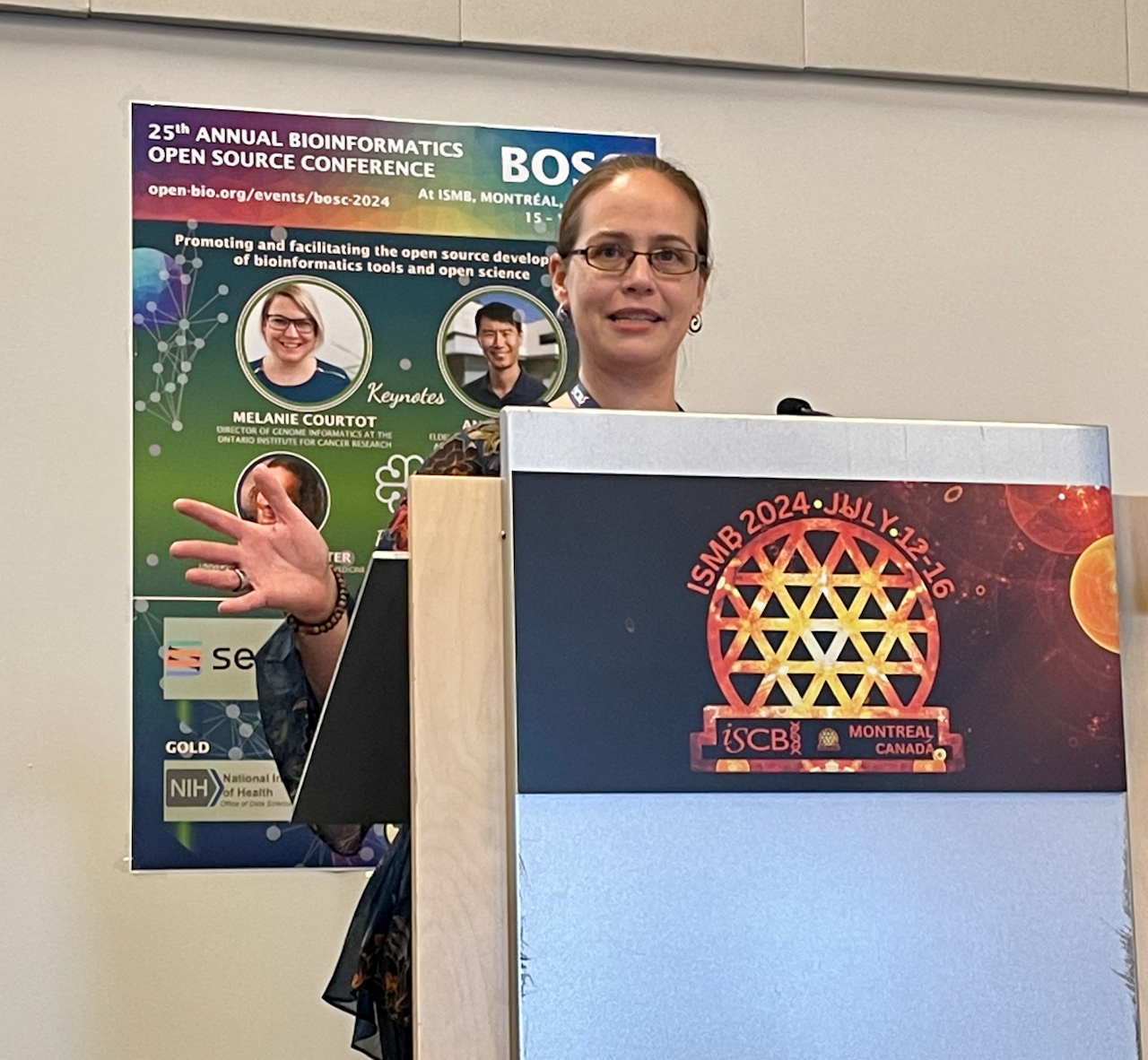


OBF Event Awards
The OBF Event Fellowship program aims to increase diverse participation at events promoting open source bioinformatics software development and open science in the biological research community.
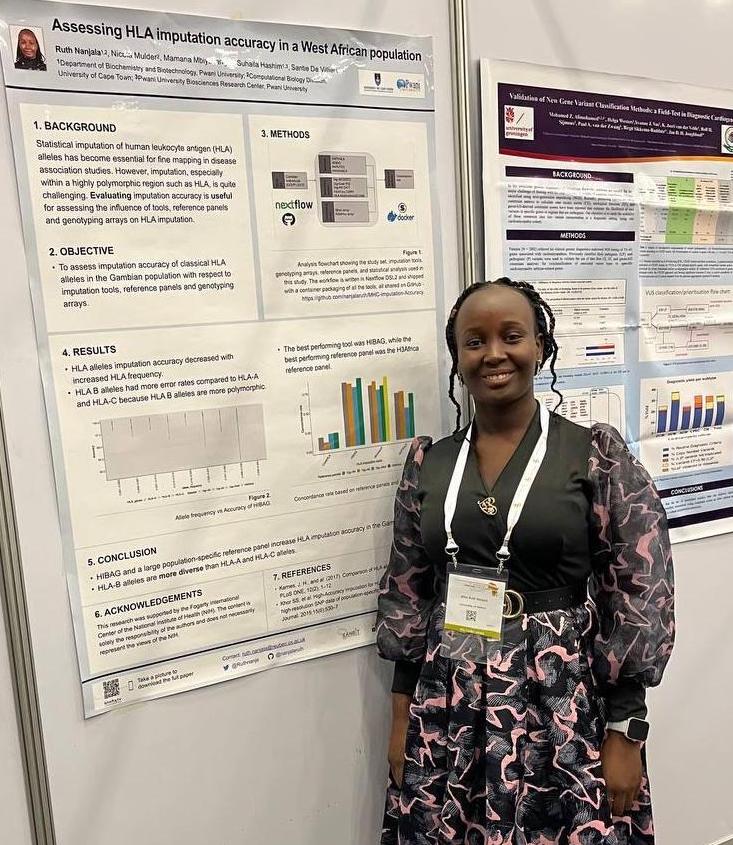
They let me into Australia, and this is what I saw
Montreal BioJava Bootcamp Announced
BioneQ, the Quebec Bioinformatics Network, is organizing the first North American BioJava Bootcamp from August 18th to 22nd. We have invited Matthew Pocock to come to Montreal to present the material that has been presented to the European Bootcamps for quite some time now. On the agenda (preliminary):
-Sequence I/O and manipulations; -BLAST and FASTA parsing; -Using databases with BioJava; -Intro to Sequence GUI.
The bootcamp will be at the Universite de Montreal and the registration fee is $250US. If you are interested, use the following link to register:
[Read More]BioPerl workshop materials online
Heikki has converted some Bioperl workshop materials to a web slide show that is available here:
BioJava 1.3 Released
Thomas Down writes:
After a long series of pre-releases (and many bug fixes), I’ve just finished building BioJava 1.30. Source, binaries, and javadocs can all be found at:
http://www.biojava.org/download/
As with the pre-releases, separate binaries are available for java platform releases 1.3 and 1.4. The 1.4 releases include some extra features which depend on jdk1.4 extensions such as the java.nio package.
Highlights of this release include:
- Packed storage of sequence data in memory
- Better support for the OBDA database access standards
- Improvements to the parsers for output from tools like blast and fasta.
- Many enhancements to the FeatureFilter system.
[Read More]Chromatogram viewing with Java
Rhett Sutphin provides a mini-primer and some example code that shows how to view Chromatogram files with java.
His full posting can be read here: http://pw600a.bioperl.org/pipermail/biojava-l/2003-June/003896.html
The Open Biological Database Access (OBDA) introduction for BioPerl
Do you need to access sequences from multiple places? Would you like to easily retrieve your own local sequences from indexed flat files, all other sequences on species X from department wide raletional database and the rest from global internet servers?
The Open Biological Database Access (OBDA) System was designed so that one could use the same application code to access data from all three of the database types by simply changing a few lines in a “configuration file”. This makes application code more portable and easier to maintain.
[Read More]BioJava tutorial updated
Mark writes:
The tutorial site BioJava in Anger has been updated so the code examples reflect the new APIs in the upcomming BioJava 1.3 release. Take a peek at http://bioconf.otago.ac.nz/biojava
bioperl-run 1.2.0 released
Shawn Hoon announces the release of bioperl-run-1.2.0 which is an extention to the bioperl framework that contains modules that act as wrappers for common informatics applications.
The full release announcement can be read here: http://bioperl.org/pipermail/bioperl-l/2003-May/012308.html
Avantgo-friendly channel for bioperl list
Jonathan Epstein has a simple script and directions for browsing bioperl list traffic offline via a mobile device.
The full post can be read online at: http://bioperl.org/pipermail/bioperl-l/2003-May/012287.html